2026 | |
| Three-State Gene Expression Model Parameterized for Single-Cell Multi-Omics Data T Peyric, T Lepoutre, A Crombach, T Guyet In: Fages, F., Pérès, S. (eds) Computational Methods in Systems Biology. CMSB 2025. Lecture Notes in Computer Science, vol 15959. Springer, Cham. (see also: bioRxiv) |
2025 | |
| Explaining Random Forest and XGBoost with Shallow Decision Trees by Co-clustering Feature Importance
RG Pensa, A Crombach, S Peignier, C Rigotti Mach Learn 114, 287 |
 | Mammalian olfactory cortex neurons retain molecular signatures of ancestral cell types
S Zeppilli, A Ortega Gurrola, P Demetci, D Brann, R Attey, N Zilkha, T Kimchi, SR Datta, R Singh, MA Tosches, A Crombach, A Fleischmann Nature Neuroscience, 28:5, 1546–1726 (preprint: bioRxiv) |
| Re_actShap: détection de rebranchements des réseaux de régulation d’expression génique à l’aide des valeurs SHAP
L Chabrier, A Crombach, S Peignier, C Rigotti 25ième conference sur l'Extraction et Gestion des Connaissances (EGC), session demonstrations |
2024 | |
| Combining SHAP-driven Co-clustering and Shallow Decision Trees to Explain XGBoost
RG Pensa, A Crombach, S Peignier, C Rigotti Proc. of 27th International Conference Discovery Science (DS 2024), 369–384 (preprint: HAL) |
| Detecting gene regulatory network rewiring as changes in marker regulatory associations using only single-cell gene expression data
L Chabrier, A Crombach, S Peignier, C Rigotti IEEE 12th Data Mining in Biomedical Informatics and Healthcare Workshop (DMBIH'24) (preprint: HAL) |
 | Effective pruning for top-k feature search on the basis of SHAP values
L Chabrier, A Crombach, S Peignier, C Rigotti IEEE Access, 12, 163079–163092 (preprint: HAL) |
2021 | |
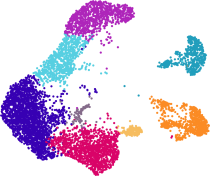 | Molecular characterization of projection neuron subtypes in the mouse olfactory bulb
S Zeppilli, T Ackels, R Attey, N Klimpert, KD Ritola, S Boeing, A Crombach, AT Schaefer, A Fleischmann eLife 10:e65445 (preprint: bioRxiv) |
2020 | |
 | The genomic view of diversification
J Marin, G Achaz, A Crombach, and A Lambert Journal of Evolutionary Biology, 33:10, 1387–1404 (preprint: bioRxiv) |
2019 | |
 | Evolution of the D. melanogaster chromatin landscape and its associated proteins
E Parey and A Crombach Genome Biology and Evolution, evz019 (preprint: bioRxiv) |
2018 | |
 | A damped oscillator imposes temporal order on posterior gap gene expression in Drosophila
B Verd, E Clark, KR Wotton, H Janssens, E Jiménez-Guri, A Crombach, and J Jaeger PLoS Biology, 16:2, e2003174 (preprint: bioRxiv) |
2017 | |
 | Scatter search applied to the inference of a development gene network
AM Abdol, D Cicin-Sain, JA Kaandorp, and A Crombach Computation, 5:2, 22 |
 | Dynamic maternal gradients control timing and shift-rates for gap gene expression
B Verd, A Crombach, and J Jaeger PLoS Computational Biology, 13:2, e1005285 (preprint: bioRxiv) |
2016 | |
 | Gap gene regulatory dynamics evolve along a genotype network
A Crombach*, KR Wotton*, E Jiménez-Guri, and J Jaeger Molecular Biology and Evolution, 33:5, 1293–1307 (preprint: bioRxiv) |
2015 | |
 | Quantitative system drift compensates for altered maternal inputs to the gap gene network of the scuttle fly Megaselia abdita
KR Wotton, E Jiménez-Guri, A Crombach, H Janssens, A Alcaine-Colet, S Lemke, U Schmidt-Ott, and J Jaeger eLife 2015;4:e04785 |
 | High-resolution gene expression data from blastoderm embryos of the scuttle fly Megaselia abdita
KR Wotton, E Jiménez-Guri, A Crombach, D Cicin-Sain, and J Jaeger Scientific Data 2, Article number 150005 |
 | SuperFly: a comparative database for quantified spatio-temporal gene expression patterns in early dipteran embryos
D Cicin-Sain*, A Hermoso Pulido*, A Crombach*, KR Wotton, E Jiménez-Guri, JF Taly, G Roma, and J Jaeger Nucleic Acids Research, 43 D751–D755 |
 | BioPreDyn-bench: a suite of benchmark problems for dynamic modelling in systems biology
AF Villaverde, D Henriques, K Smallbone, S Bongard, J Schmid, D Cicin-Sain, A Crombach, J Saez-Rodriguez, K Mauch, E Balsa-Canto, P Mendes, J Jaeger and JR Banga BMC Systems Biology, 9:8 |
2014 | |
 | Classification of transient behaviours in a time-dependent toggle switch model
B Verd, A Crombach, and J Jaeger BMC Systems Biology, 8:1, 43 |
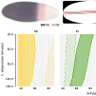 | Evolution of early development in dipterans: reverse-engineering the gap gene network in the moth midge Clogmia albipunctata (Psychodidae)
A Crombach, M Garcia-Solache, and J Jaeger Biosystems, 123 74–85 |
2013 | |
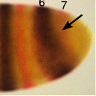 | Lack of tailless leads to an increase in expression variability in Drosophila embryos
H Janssens, A Crombach, KR Wotton, D Cicin-Sain, S Surkova, LC Lu, M Samsonova, M Akam, and J Jaeger Developmental Biology, 377:1, 305–317 |
2012 | |
 | Efficient reverse-engineering of a developmental gene regulatory network
A Crombach*, KR Wotton*, D Cicin-Sain, M Ashyraliyev, and J Jaeger PLoS Computational Biology, 8:7, e1002589 |
 | Medium-throughput processing of whole mount in situ hybridisation experiments into gene expression domains
A Crombach*, D Cicin-Sain*, KR Wotton*, and J Jaeger PLoS One, 7:9, e46658, 2012 |
2011 | |
 | Is RNA-dependent RNA polymerase essential for transposon control?
A Crombach and P Hogeweg BMC Systems Biology, 5:1, 104 |
2009 | |
 | Evolution of resource cycling in ecosystems and individuals
A Crombach and P Hogeweg BMC Evolutionary Biology, 9:1, 122 |
2008 | |
 | Evolution of evolvability in gene regulatory networks
A Crombach and P Hogeweg PLoS Computational Biology, 4:7, e1000112 |
2007 | |
 | Chromosome rearrangements and the evolution of genome structuring and adaptability
A Crombach and P Hogeweg Molecular Biology and Evolution, 24:5, 1130–1139 |