Last May, I wrote that I would explore Scala for prototyping and potential outreach activities. Over the last half year, I have been reading its on-line documentation, relevant blogs and books, and example code on Github; and I have tried writing some code snippets. I encountered a bit of a learning curve —especially when it come to Scala’s type system acrobatics— but, all in all, the language is rather pragmatic. Simple tasks are simple to implement, difficult ones a bit less simple. In summary, the moment has come to share a first few bits of code.
Before I present three little programs, I would like to thank Darren Wilkinson (https://darrenjw.github.io/). His work was instrumental to put the right ideas in my head and I appreciate his rationale that for developing stats, data science, and simulation code, we should take advantage of modern languages such as Scala (or Kotlin, Rust) over older ones like C/C++, Fortran, R, Matlab, and so on.
Now, for my first adventures in Scala I went back to the “Bioinformatic Processes” course of Paulien Hogeweg that I followed in the early 2000s. That course had some beautiful examples of spatial patterning and I was keen to recreate them. I dug up the simulation toolkit used in the course, called Cellular Automata in Simulated Hardware (CASH), and adapted a tiny part of its interface to my needs. I figured a tiny bit of cash is a few cents, hence the name of my Github repository: scala-cents.
In the Github repository and in the screenshots below, you can find:
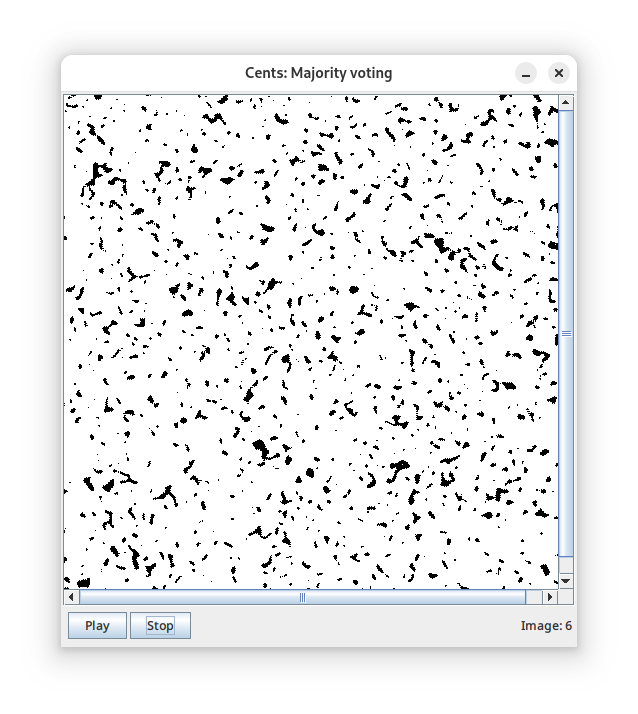
- A simple test case: a voting rule cellular automaton (left panel);
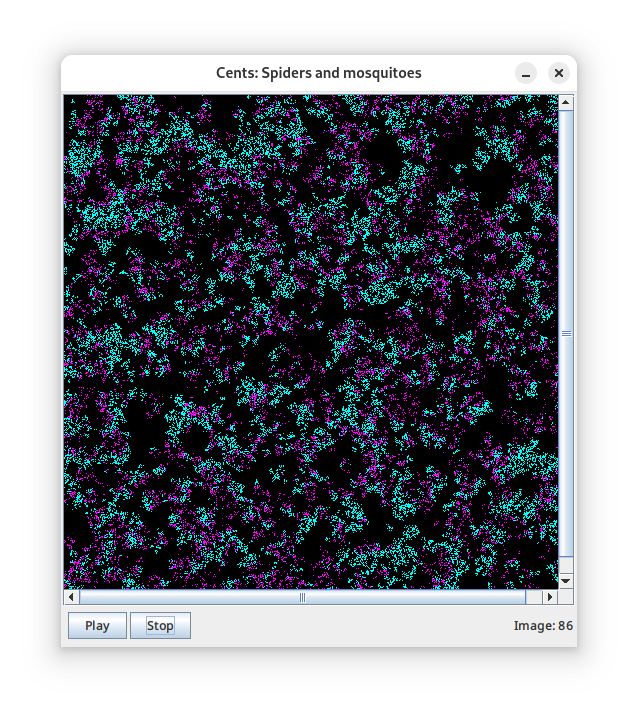
- A predator-prey simulation of spiders that eat mosquitoes. Some settings give pretty, “turbulent” wave patterns (middle);
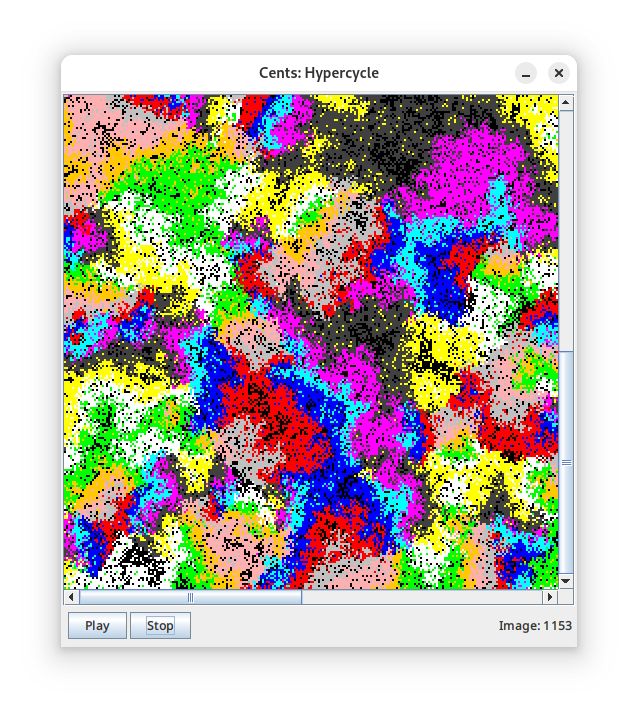
- And a classic origin-of-life scenario called the hypercycle. In the hypercycle, each bio-molecule catalyzes the copying of the next bio-molecule in the cycle. If you define a cycle > 3, you get spirals (right).
It should not surprise anyone that my next step will be to design and code support for gene regulatory networks.